Apps and Visualisation
The Apps module in Quark V3 allows users to create and configure custom visualisation applications for exploring and interpreting pipeline outputs. Apps are containerised tools that run within the platform, giving researchers an interactive view of their data without leaving Quark.
Audience: Bench scientists and bioinformaticians
Access: Select Apps from the left navigation pane
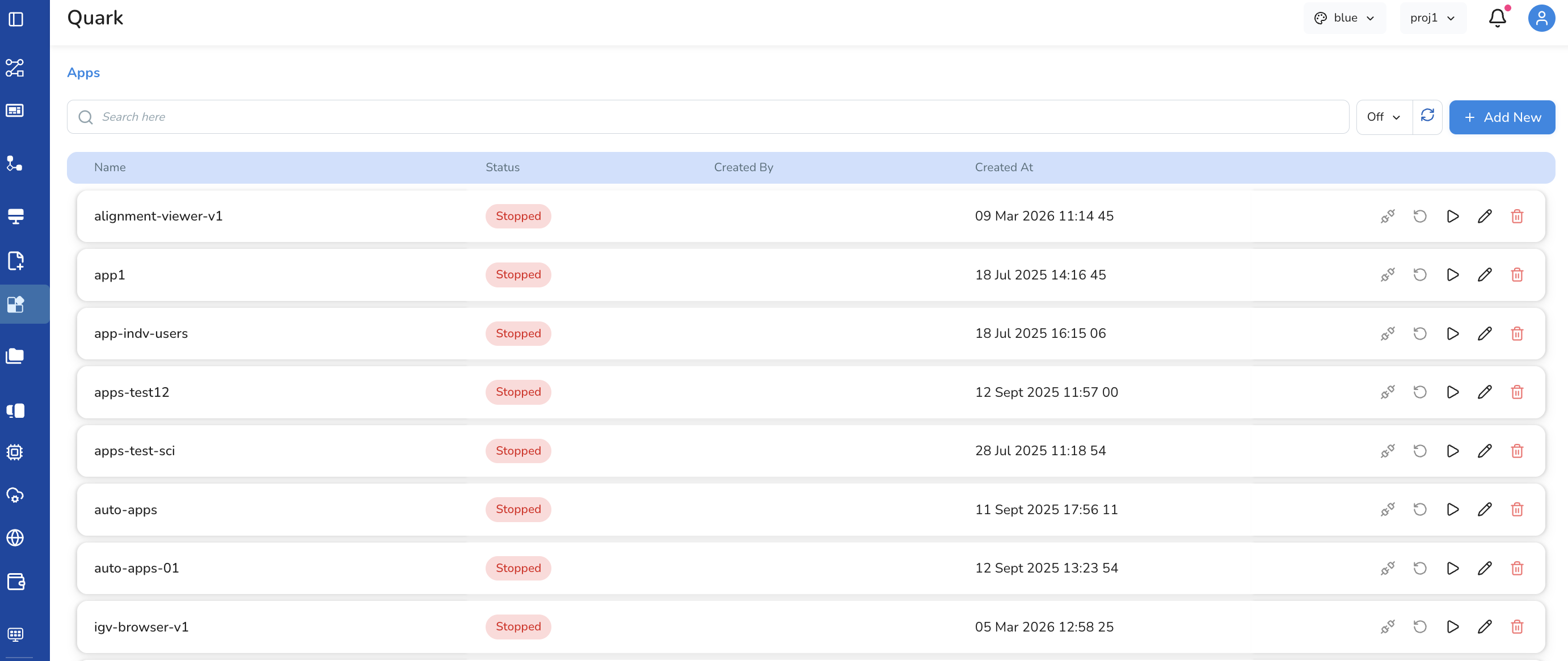
Overview
Apps in Quark V3 are fully user-configured. Each app is set up by specifying a container image, compute resources, dataset mounts, and permissions. Once configured, an app can be connected, re-run, started, edited, or deleted.
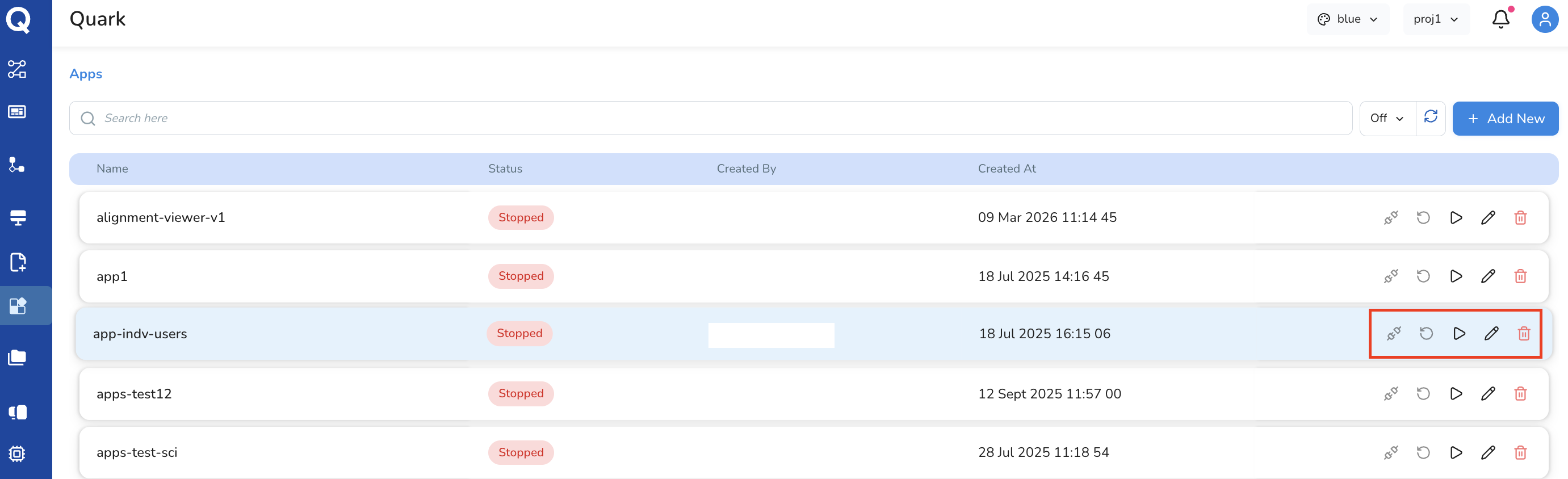
Each app row in the Apps list has the following action icons:
| Icon | Action |
|---|---|
| Connect | Start and connect to the app |
| Re-run | Re-launch the app with the same configuration |
| Start | Start a previously stopped app |
| Edit | Modify the app's configuration |
| Delete | Permanently remove the app |
Creating a New App
To add a new visualisation app:
- Select Apps from the left navigation pane
- Click Add New in the top right corner of the display screen
- Complete the four configuration tabs: General, Services, Datasets, and Permissions
Tab 1: General
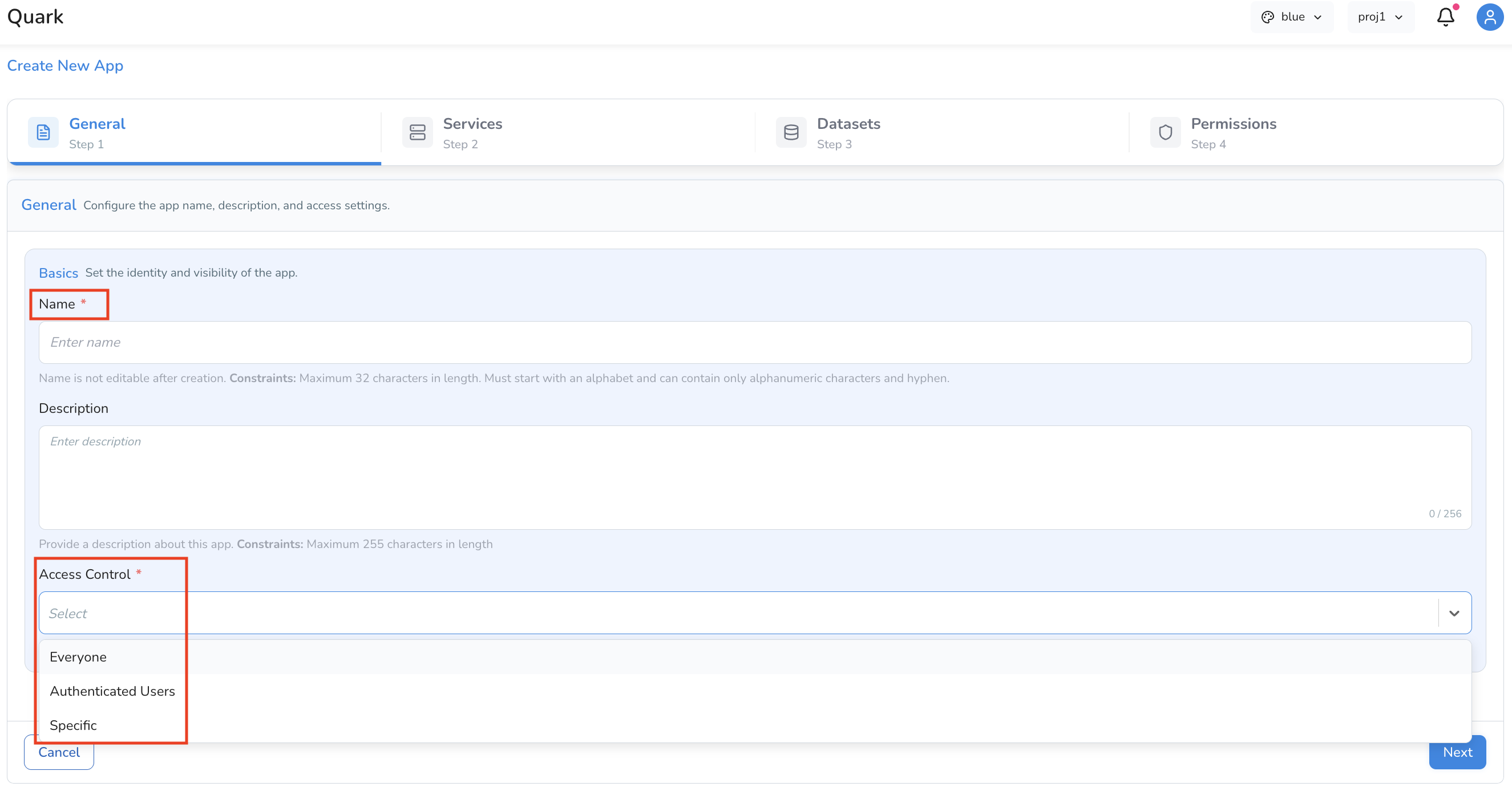
Provide the basic metadata for your app:
| Field | Description |
|---|---|
| Name | A short, descriptive name for the app (e.g. IGV-peptide-viewer) |
| Description | A brief summary of the app's purpose |
| Access Control | Select who can access this app: Everyone, Authenticated Users, Specific |
Tab 2: Services
The Services tab is where you define the container image, compute resources, and runtime configuration for your app. Each app requires at least one service, and you can add multiple services by clicking Add New Service at the bottom of the tab.
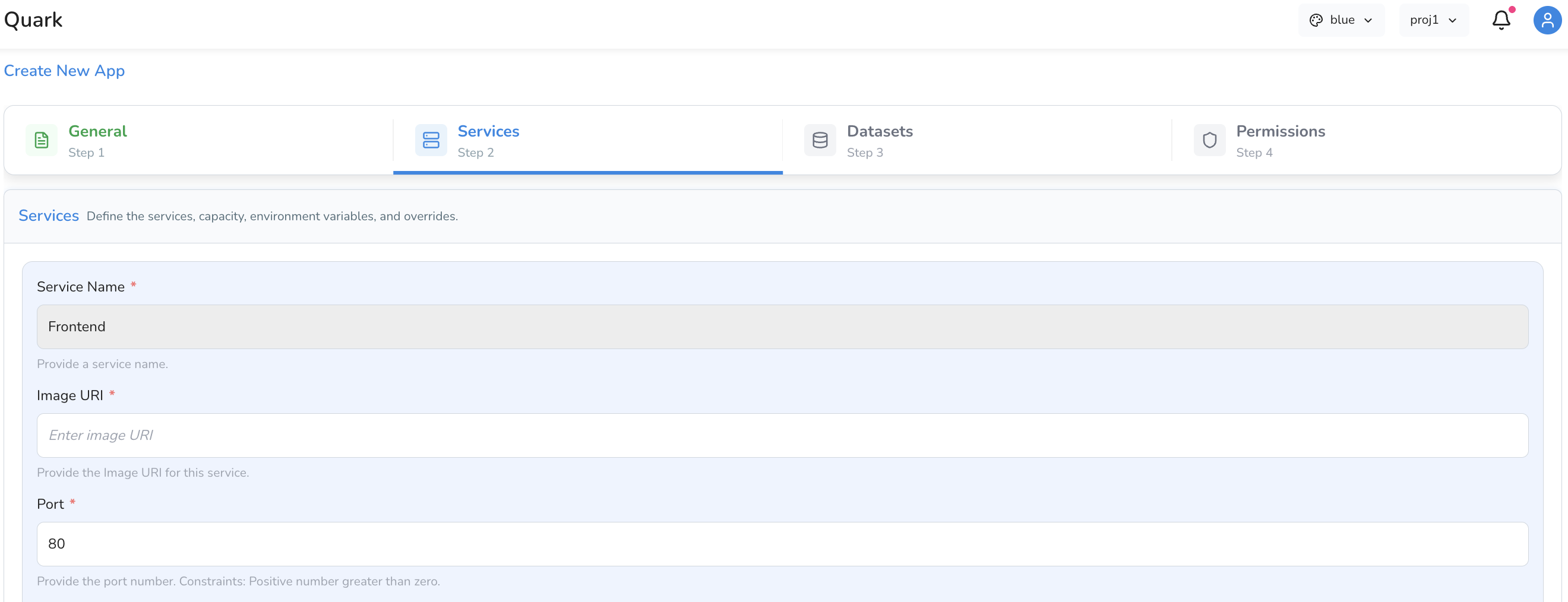
Service Fields
Identity and Container
| Field | Description |
|---|---|
| Service Name | A name for this service. Defaults to Frontend. Update this if you are adding multiple services to distinguish them. |
| Image URI | The container image URI for the visualisation tool — for example, a Docker image hosted on AWS ECR or Docker Hub. |
| Port | The port the service will listen on. Defaults to 80. Update this if your container image exposes a different port. |
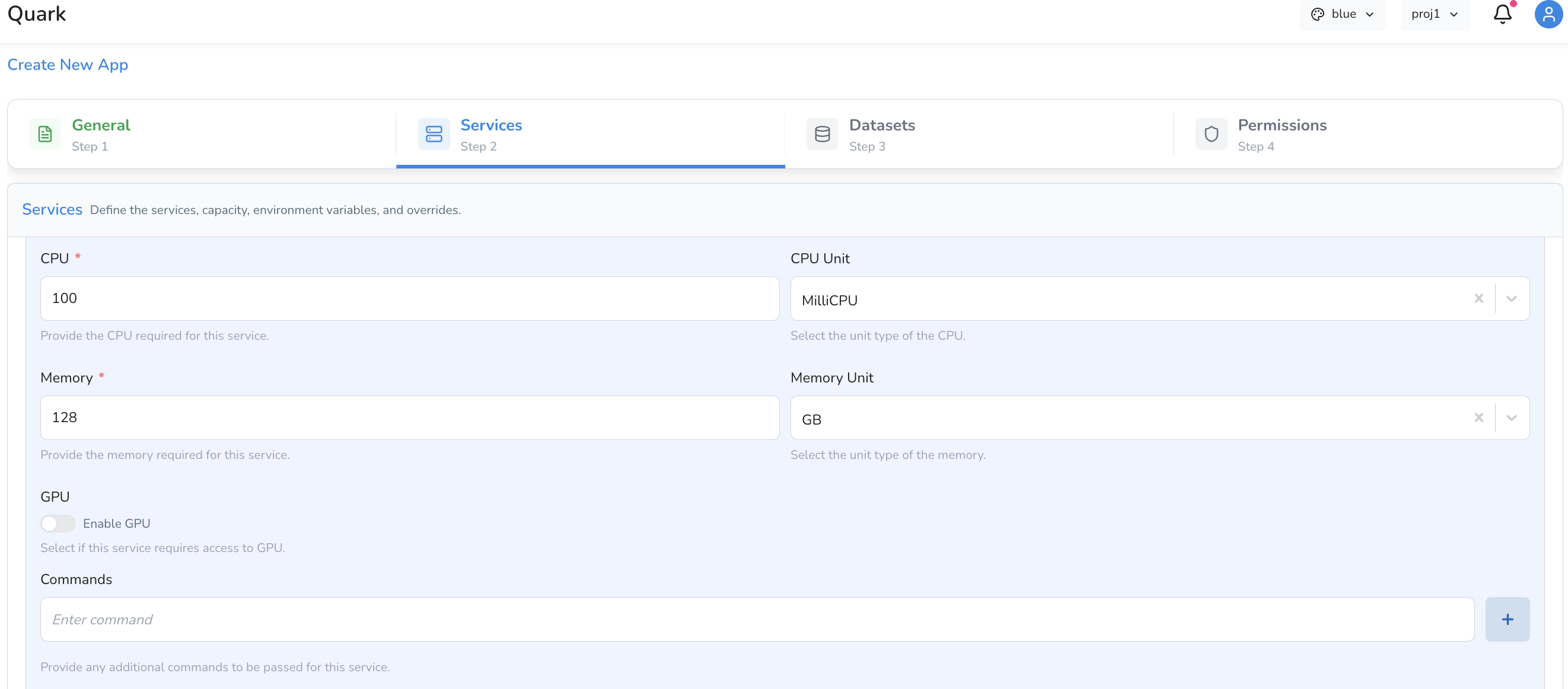
Compute Resources
| Field | Description |
|---|---|
| CPU | The number of CPU units to allocate to this service. |
| CPU Unit | The unit for the CPU value (e.g. millicores or cores, depending on your platform configuration). |
| Memory | The amount of memory to allocate to this service. |
| Memory Unit | The unit for the memory value (e.g. Mi for mebibytes, Gi for gibibytes). |
| GPU | Toggle on if the service requires GPU acceleration. |
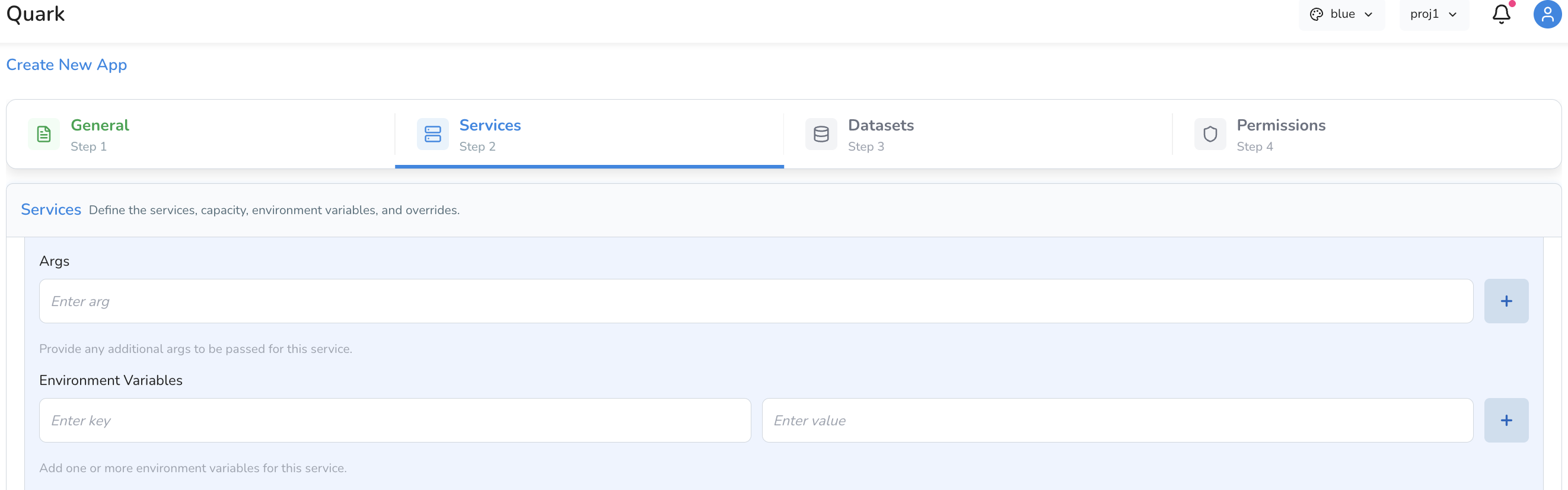
Runtime Configuration
| Field | Description |
|---|---|
| Commands | The command(s) to run when the container starts. Overrides the default CMD or ENTRYPOINT defined in the container image. Leave blank to use the image default. |
| Args | Arguments to pass to the container's entry command. Leave blank if not required. |
| Environment Variables | Key-value pairs made available as environment variables within the running container. Useful for passing configuration values such as file paths, API endpoints, or feature flags. |
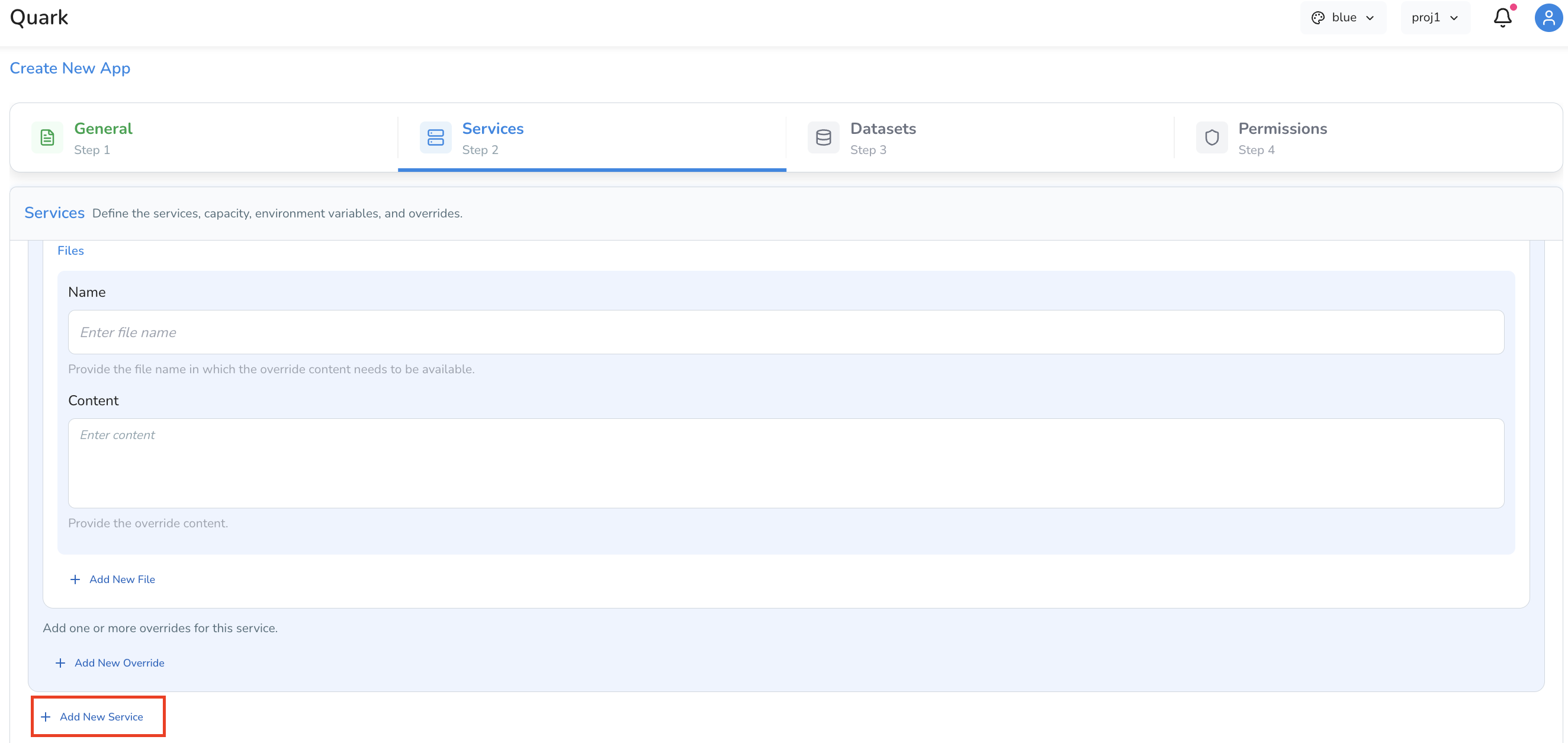
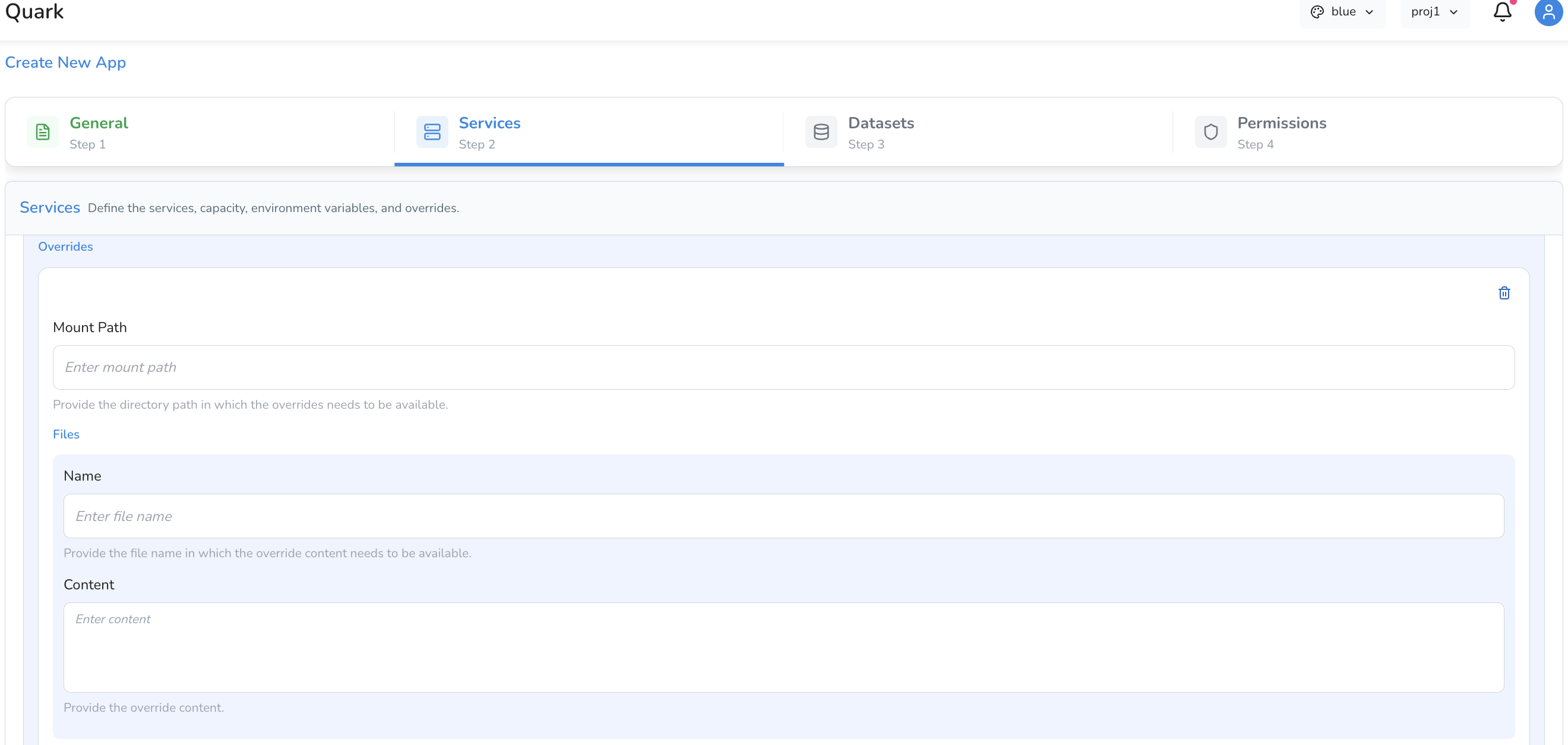
Overrides
Overrides allow you to mount configuration files directly into the container at a specified path — useful for supplying config files, reference files, or scripts that the app needs at runtime.
Each override entry has the following fields:
| Field | Description |
|---|---|
| Mount Path | The file path inside the container where the file will be mounted (e.g. /app/config/settings.json). |
| File Name | The name of the file to be created at the mount path. |
| Content | The content of the file, entered directly in the configuration screen. |
You can add multiple override entries per service by clicking Add Override. This allows you to mount several configuration files into the same container.
Adding multiple services: If your app requires more than one container — for example, a frontend visualisation service and a backend data processing service — click Add New Service to configure each one independently with its own image, compute, and runtime settings.
Example: To configure an IGV (Integrative Genomics Viewer) app, set the Service Name to
igv, provide the IGV container image URI, set the Port to80, and allocate sufficient CPU and memory for the size of data you intend to visualise.
Tab 3: Datasets
The Datasets tab is where you connect your app to the data it will visualise. Apps do not connect directly to My Files → Results automatically — you must explicitly mount the relevant data here.
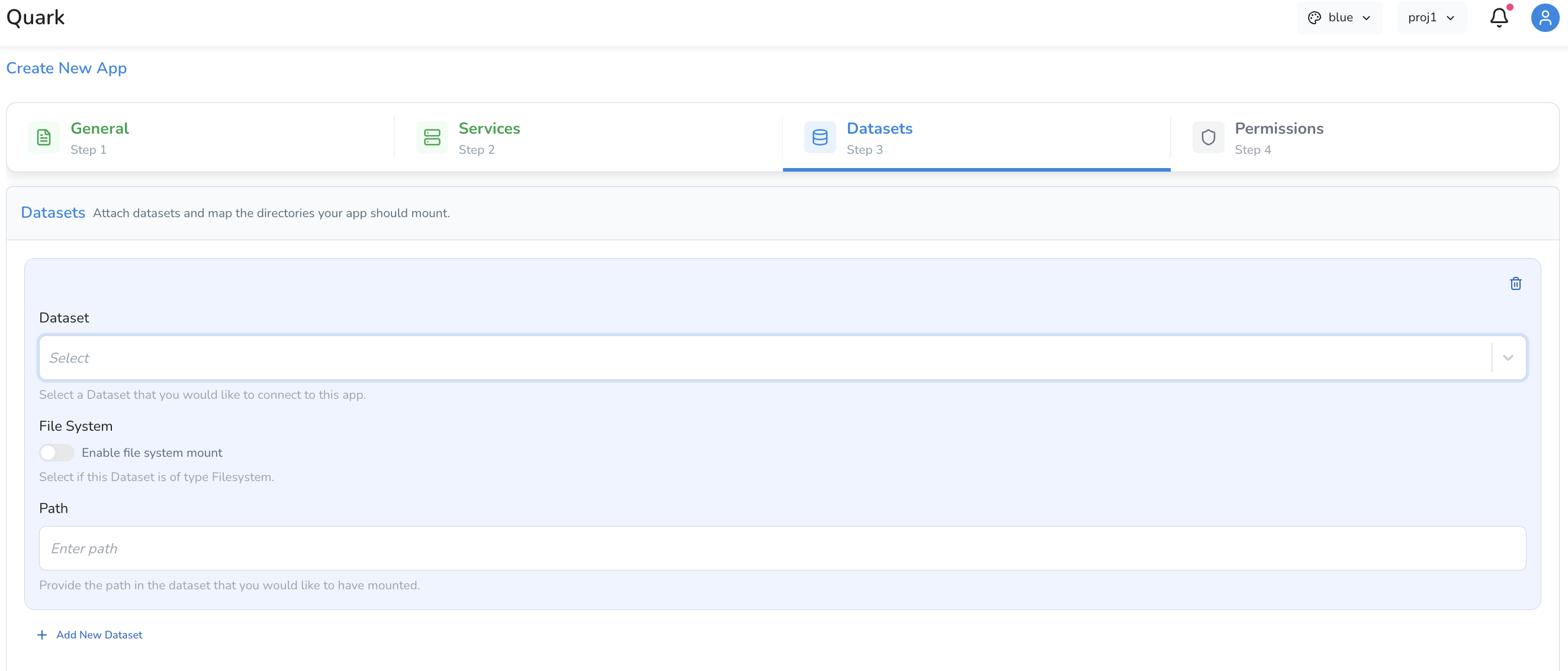
You have two options:
| Option | Description |
|---|---|
| Mount a dataset | Mount a specific dataset available on the platform directly into the app's container |
| Map a directory | Specify a directory path to mount into the app — useful for pointing to a Results directory from a completed pipeline run |
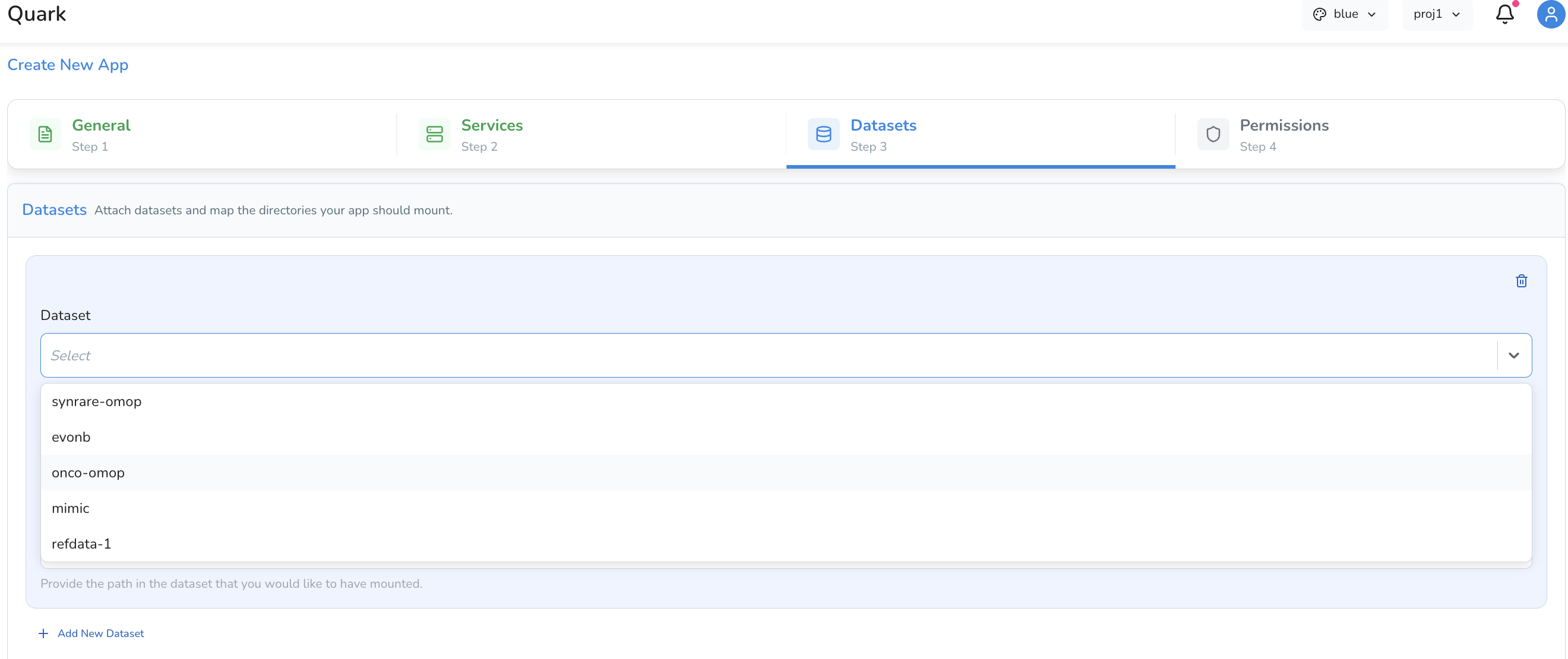
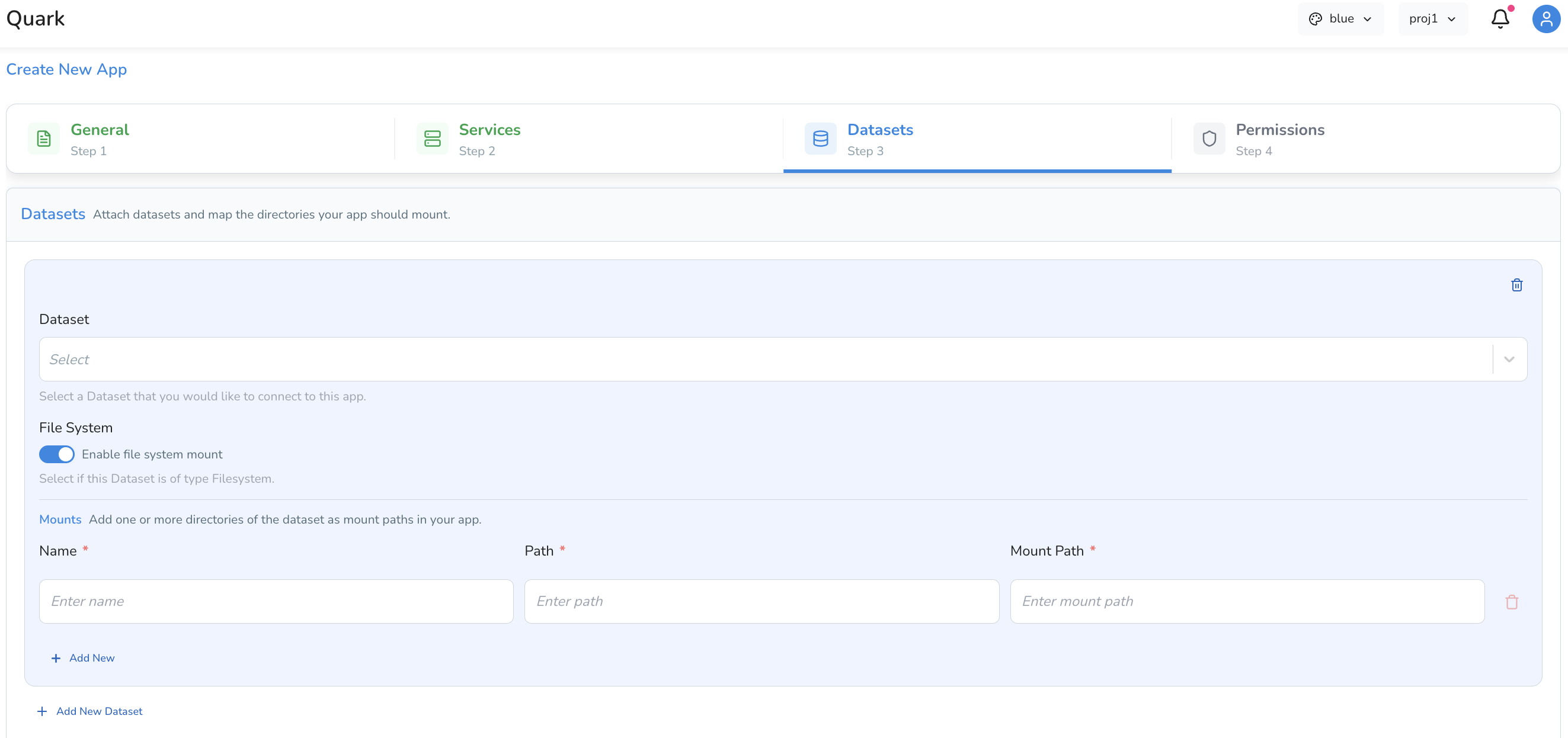
Click Add New Dataset to mount additional datasets to the app.
Tip: To visualise results from a pipeline run, use Map a directory and provide the path to the relevant output directory from My Files → Results.
Tab 4: Permissions
The Permissions tab defines who can access the app:
- Set access controls to restrict the app to yourself or share it with other members of your project or organisation
- Permissions are managed at the app level, independently of project-level access controls
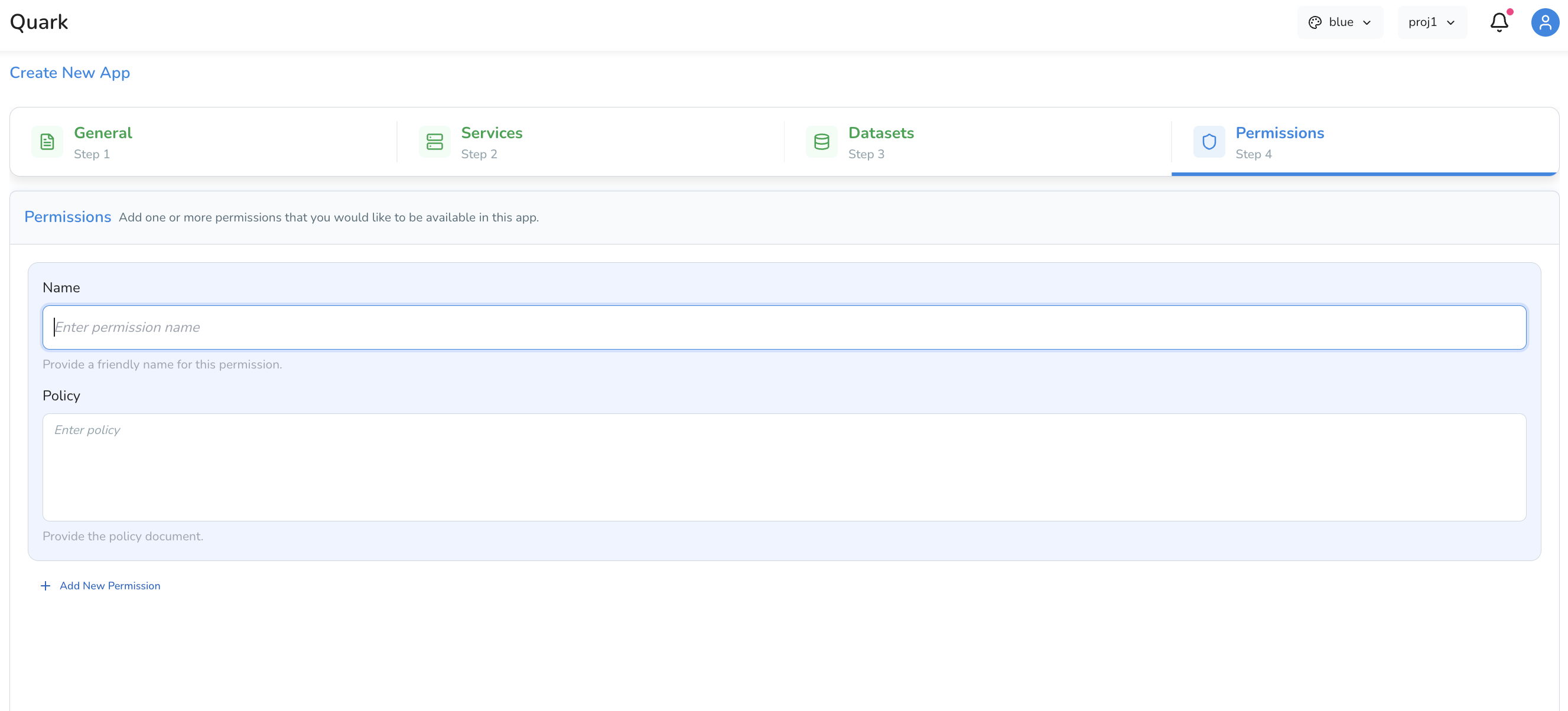
Click on the Review button to review all your app settings. Once completed to your satisfaction, click Create to bring your app to the main Apps page.
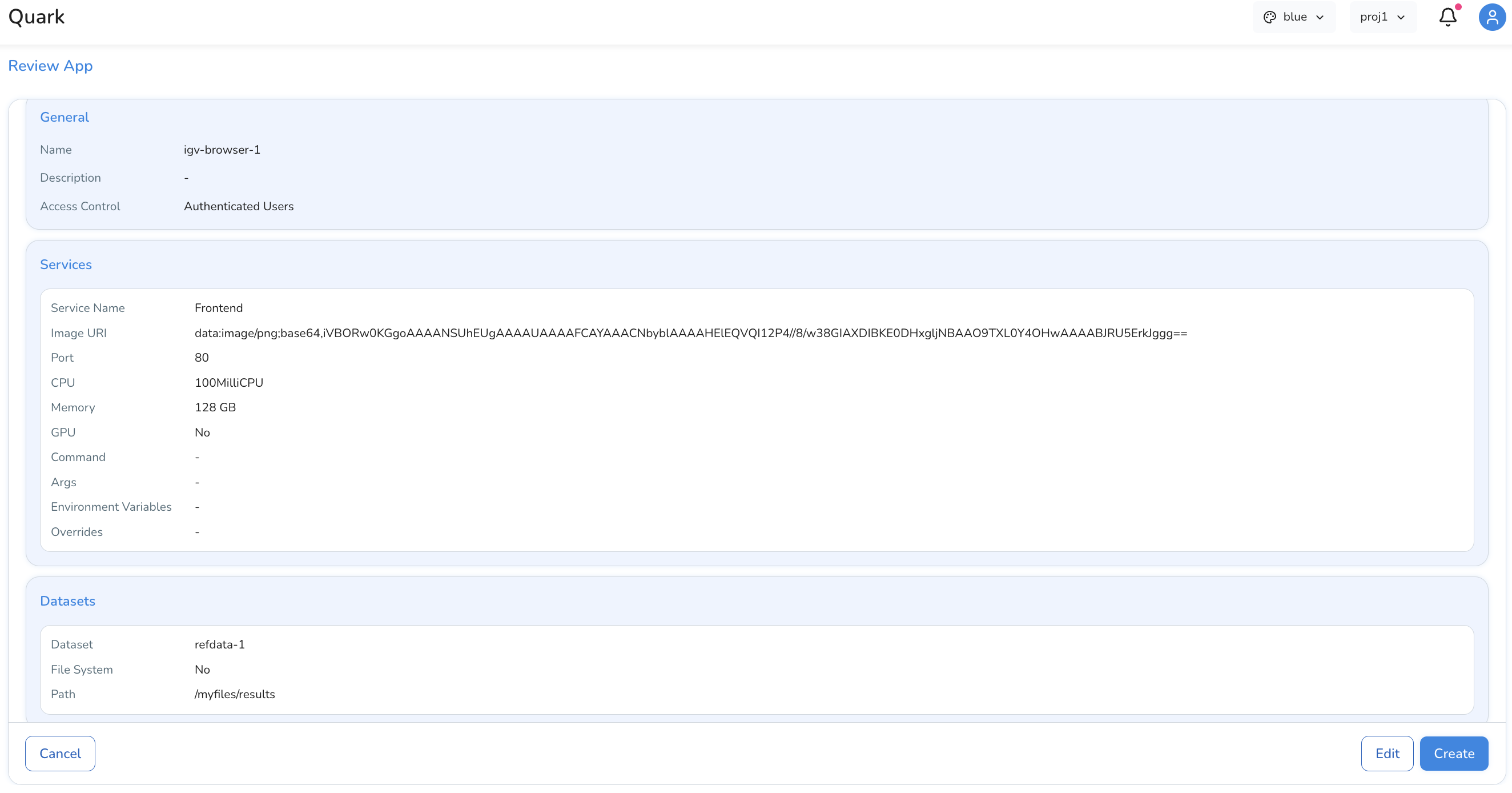
Managing Your Apps
After creation, apps are listed in the Apps tab. From here you can:
- Connect — launch and open the app
- Re-run — restart the app using its existing configuration
- Start — start a stopped app without reconfiguring it
- Edit — update any configuration tab (General, Services, Datasets, or Permissions)
- Delete — permanently remove the app and its configuration
Example: Configuring an IGV App
IGV (Integrative Genomics Viewer) is a widely used tool for visualising genomic data such as alignment files (BAM), variant call files (VCF), and genome annotations. The following is an example configuration:
General
| Field | Example value |
|---|---|
| Name | igv-dna-seq-viewer |
| Description | IGV app for reviewing DNA-Seq alignment outputs |
Services
| Field | Example value |
|---|---|
| Service Name | Frontend |
| Image URI | (your IGV container image URI from your registry) |
| Port | 80 |
| CPU | 2 |
| CPU Unit | (per your platform configuration) |
| Memory | 8 |
| Memory Unit | Gi |
| GPU | Off |
| Commands | (leave blank to use image default) |
| Args | (leave blank if not required) |
| Environment Variables | (add any required config values) |
| Overrides | (add if custom config files are needed) |
Datasets
| Field | Example value |
|---|---|
| Dataset | Map directory → path to your DNA-Seq run output in My Files → Results |
Permissions
| Field | Example value |
|---|---|
| Access | Restricted to project members |
Further Reading
- Managing Files — locating Results directories to mount in your app
- Managing Projects — add and organise workspaces and apps into projects