HealthOmics Pipelines
Quark V3 integrates natively with AWS HealthOmics — Amazon's managed service for storing, querying, and analysing genomics and biological data at scale. This integration allows bioinformaticians to import and run HealthOmics workflows directly from Quark without leaving the platform.
Audience: Bioinformaticians
Access: Select HealthOmics Pipelines from the left navigation pane
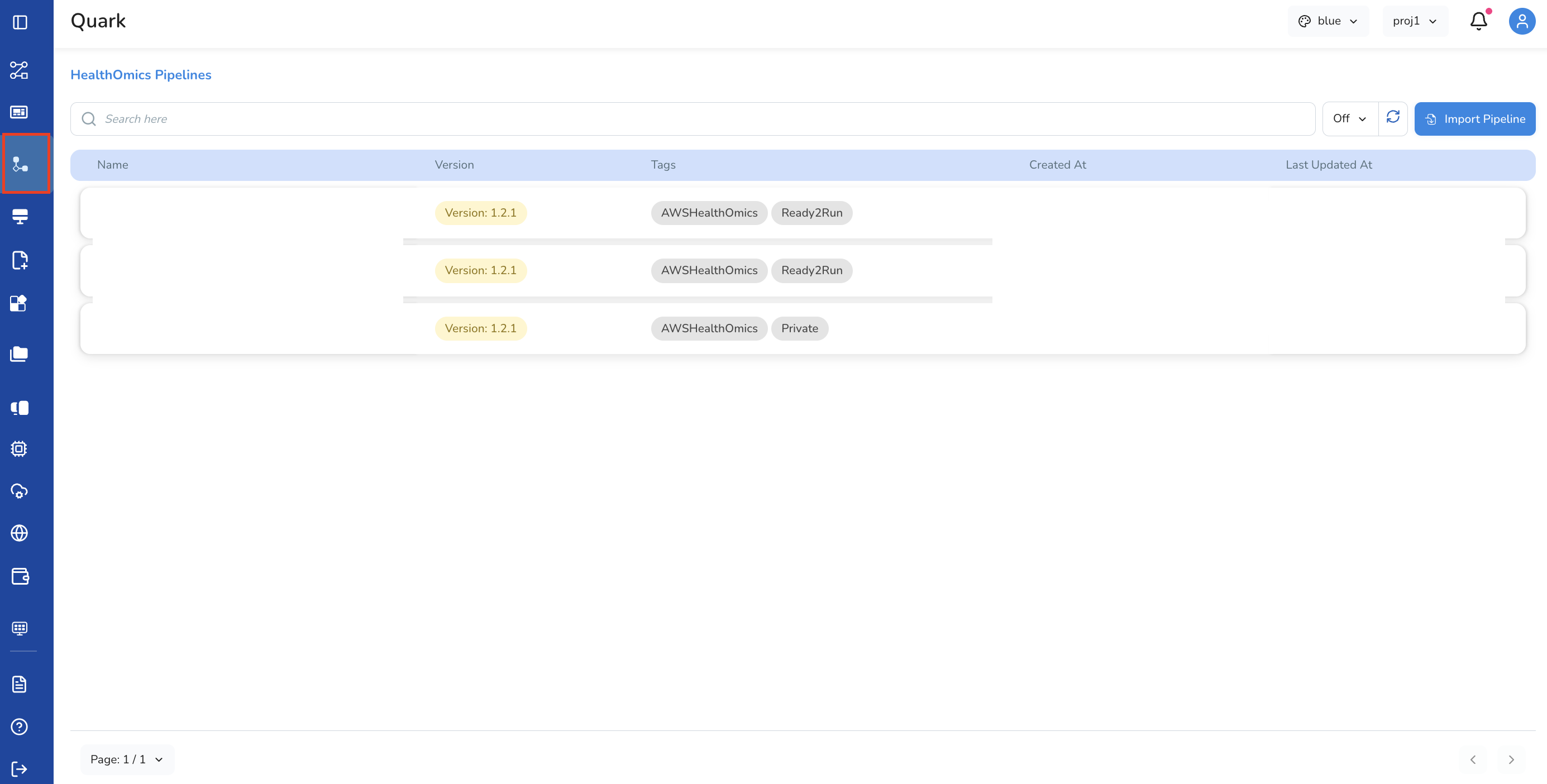
Overview
AWS HealthOmics offers two types of workflows, both accessible from Quark V3:
| Workflow Type | Description |
|---|---|
| AWS HealthOmics Private Workflow | Your organisation's own private, custom workflows — imported and run within a secure HealthOmics environment |
| AWS HealthOmics Ready2Run Workflow | Pre-validated, production-ready genomics workflows provided and maintained by AWS — available to run immediately |
Importing a Pipeline
To import a HealthOmics pipeline into Quark V3:
- Select HealthOmics Pipelines from the left navigation pane
- Click Import Pipeline
- Choose the workflow type from the two options:
- AWS HealthOmics Private Workflow
- AWS HealthOmics Ready2Run Workflow
-
Follow the import configuration steps for your chosen workflow type
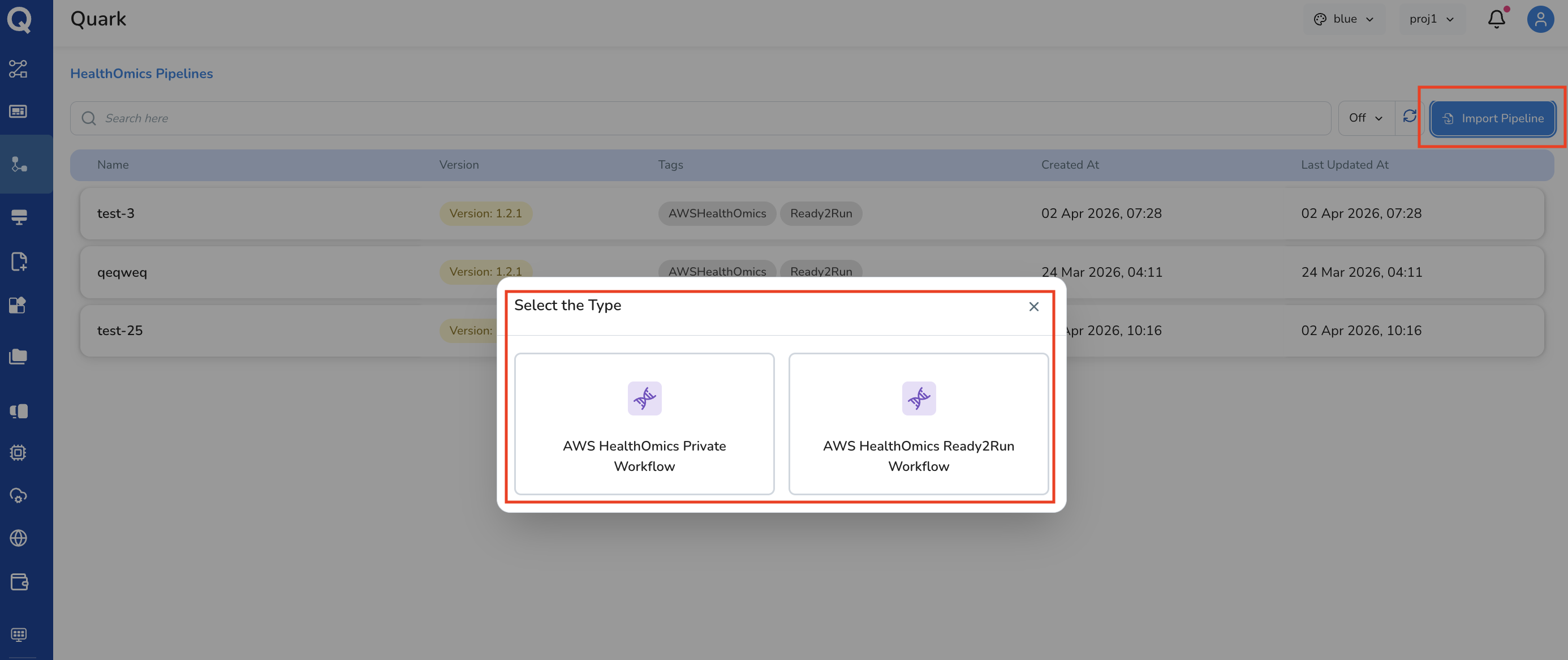
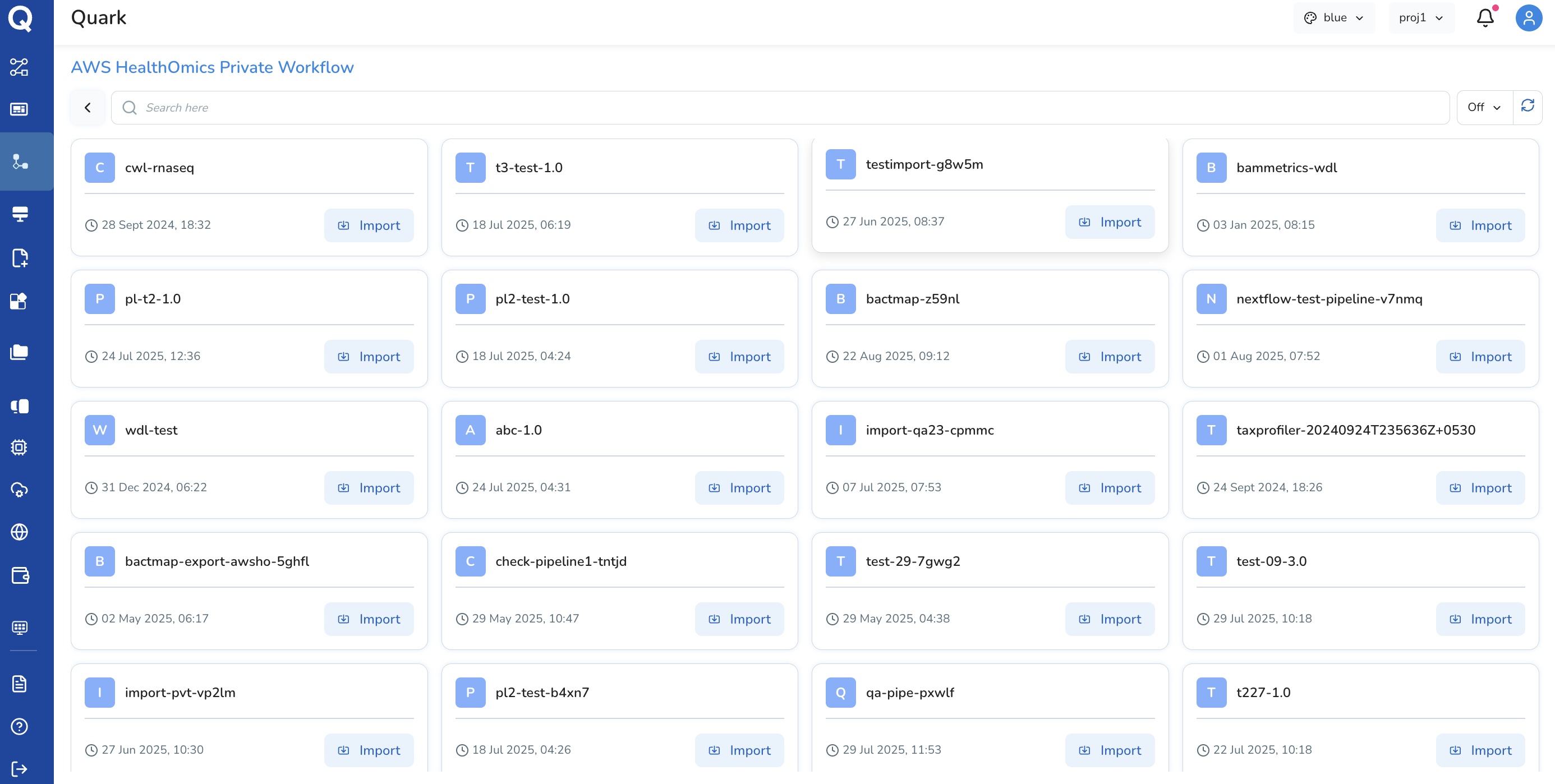
AWS HealthOmics Private Workflow
A Private Workflow is a custom pipeline your organisation has developed and stored within AWS HealthOmics. These are typically proprietary workflows built specifically for your organisation's data, processes, or research requirements.
When to use a Private Workflow: - Your organisation has existing HealthOmics workflows already deployed in AWS - You need to run proprietary, internally-developed pipelines at scale - Your data governance requirements mandate a private, organisation-owned compute environment
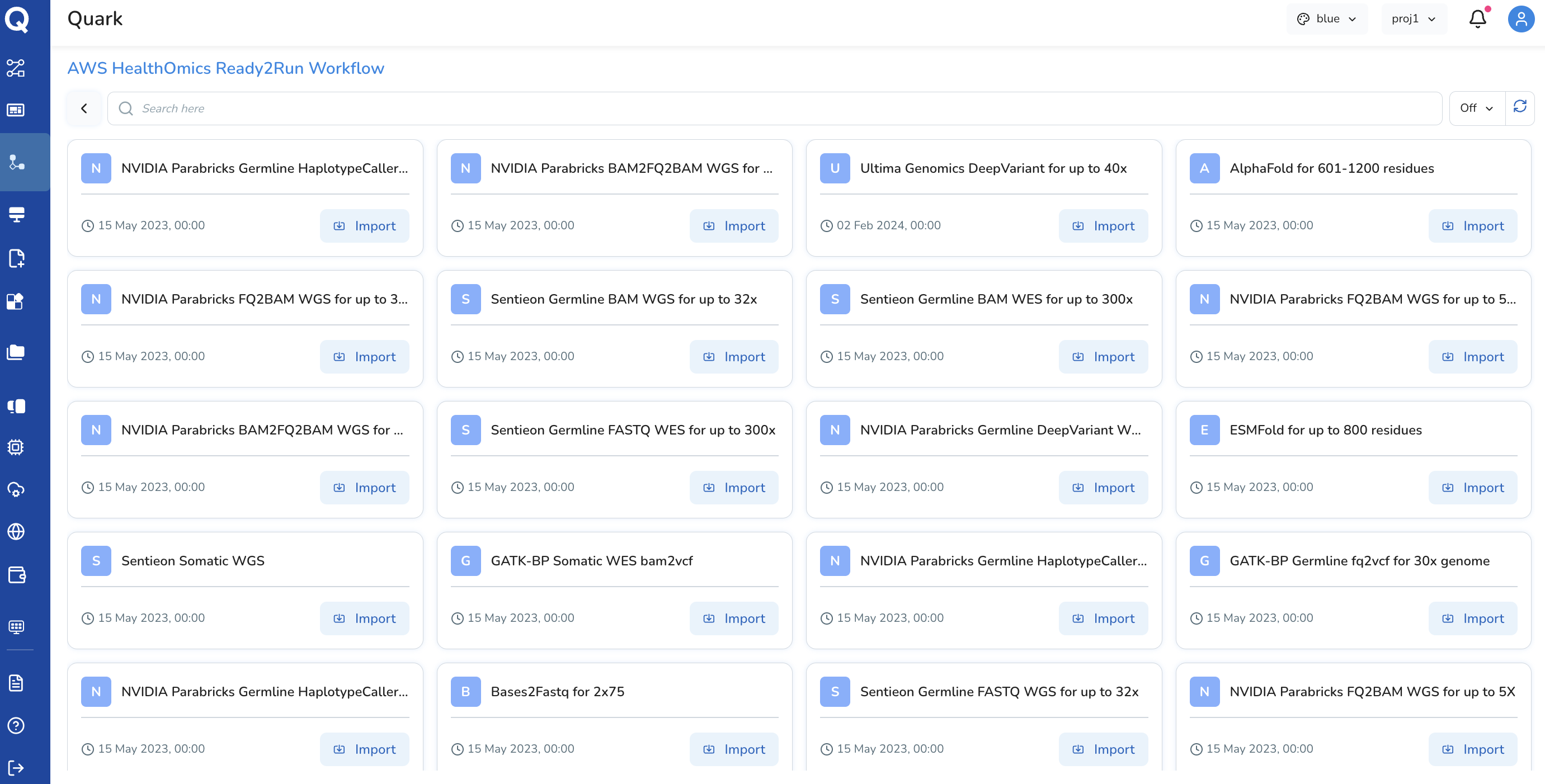
AWS HealthOmics Ready2Run Workflow
A Ready2Run Workflow is a pre-validated, curated genomics pipeline provided directly by AWS. These workflows are designed to run immediately without requiring custom setup — covering common genomics use cases such as secondary analysis of whole genome sequencing (WGS) and whole exome sequencing (WES) data.
When to use a Ready2Run Workflow: - You need a well-validated, standard genomics pipeline for a common use case - You do not have an existing custom workflow to import - You want the reliability and AWS-maintained quality assurance of a production-grade workflow
Further Reading
- AWS HealthOmics documentation — official AWS reference for HealthOmics service details