Managing Files
My Files is the central file management area in Quark V3. It is the place to upload input data before a pipeline run, access outputs after a run completes, and review a log of file activity across the platform.
Overview of Tabs
My Files is organised into three tabs:
| Tab | Purpose |
|---|---|
| Data | Upload and manage input files for use in pipeline runs |
| Results | Access output files and artefacts from completed pipeline runs |
| Activity | View a log of file-level actions and changes |
Data Tab
The Data tab is where you store input files that pipelines will use as their source data. These might include files such as:
- Protein structure files (
.pdb) - Sequencing data (
.fastq,.fastq.gz) - Alignment files (
.bam,.sam) - Reference genomes or annotation files
Uploading Files
- Navigate to My Files from the left navigation pane
- Click the Data tab
-
Click the Action button, and from the drop-down, select between New Folder (to create folders for your different projects), Upload Data, Upload from Server and CLI Auth Token
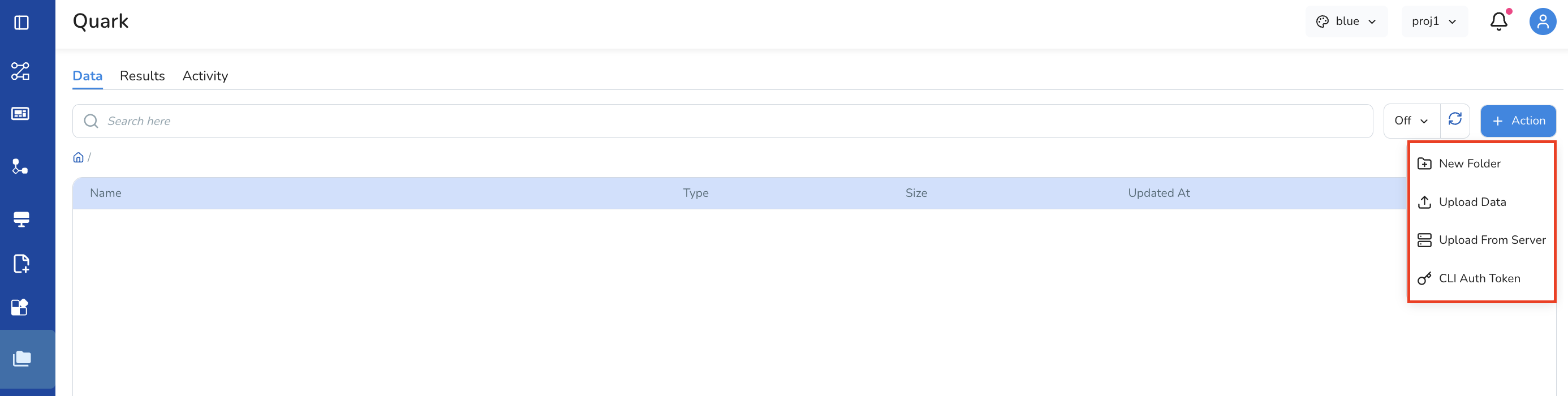
-
If you select Upload Data, select the file(s) you wish to upload from your local machine. Wait for the upload to complete — uploaded files will appear in the file list once approved by your project administrator
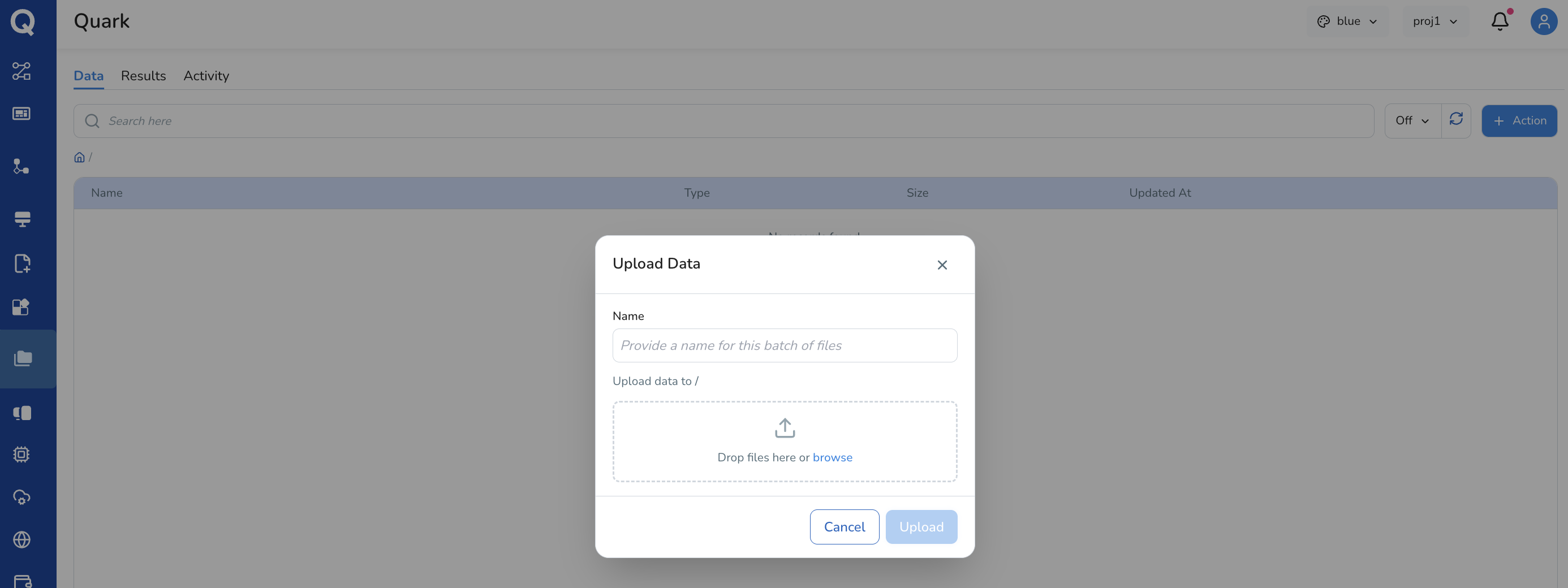
-
If you select Upload from Server, specify the Server Type (HTTP, SFTP, S3, SRA/ENA/DDBJ/GEO) and upload your data from the server location
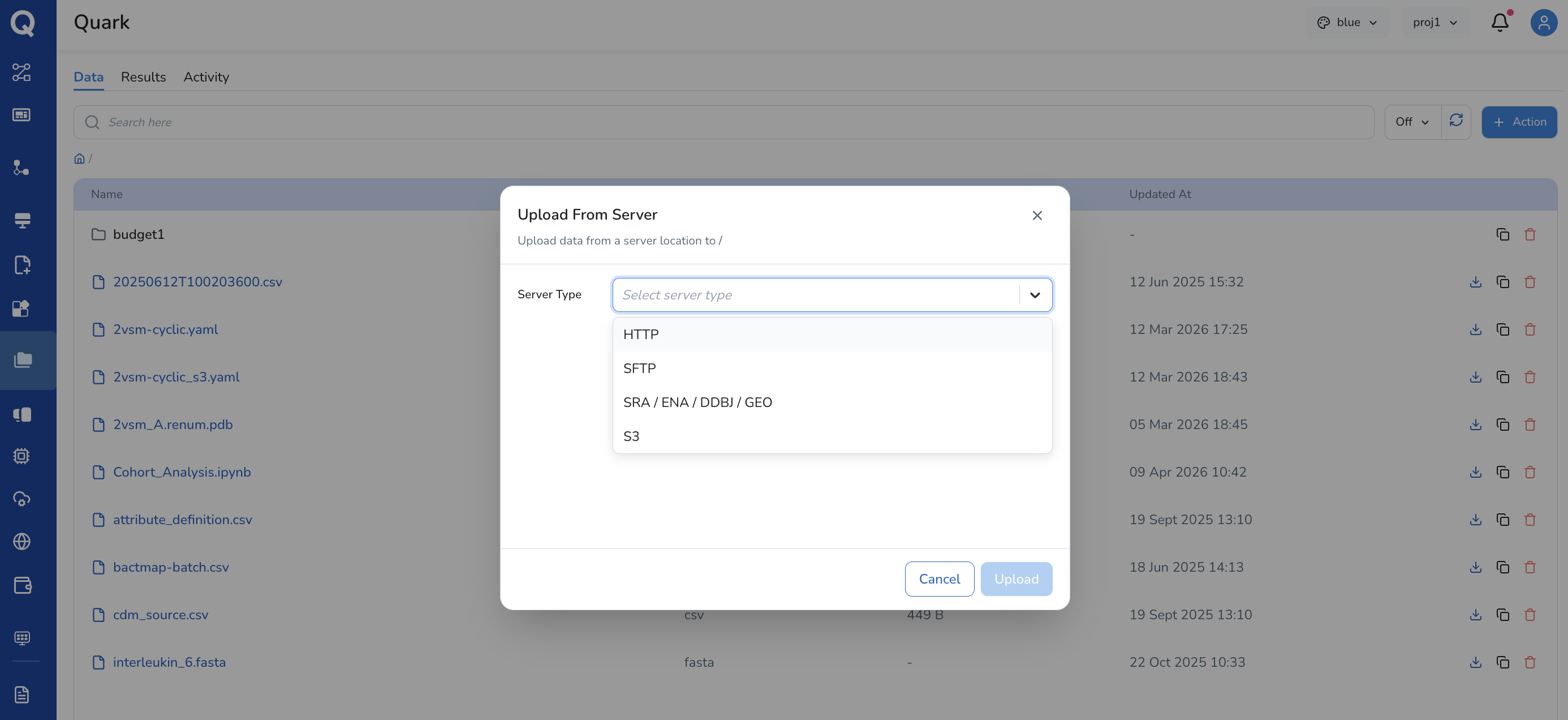
Important: For uploading and downloading large files or folders, refer to MyFiles CLI for a step-by-step guide on using the MyFiles CLI tool.
Tip: Upload your input files before configuring a pipeline run. When you reach the Inputs step during a run, you will be able to select files directly from My Files → Data.
Selecting Files in a Pipeline Run
When configuring pipeline inputs, file fields (e.g. the PDB file selector in the Boltzgen pipeline) will allow you to browse and select from files already uploaded to the Data tab. You do not need to re-upload files for each run — previously uploaded files remain available.
Results Tab
The Results tab stores all output files and artefacts generated by completed pipeline runs. Results are organised by run, making it easy to locate outputs from a specific execution.
Accessing Results
- Navigate to My Files → Results
- Browse or search for the run whose outputs you need
- Click on a result entry to view or download the associated output files
What Results Are Generated?
The outputs available in the Results tab depend on the pipeline that was run. Examples include:
- Boltzgen — designed peptide sequences, structure prediction outputs
- DNA-Seq pipelines — variant call files (VCF), alignment summaries
Refer to the documentation for your specific pipeline to understand what outputs to expect and how to interpret them.
Transferring large files: For large files or folders, use the MyFiles CLI tool. See Transferring Large Files with the MyFiles CLI.
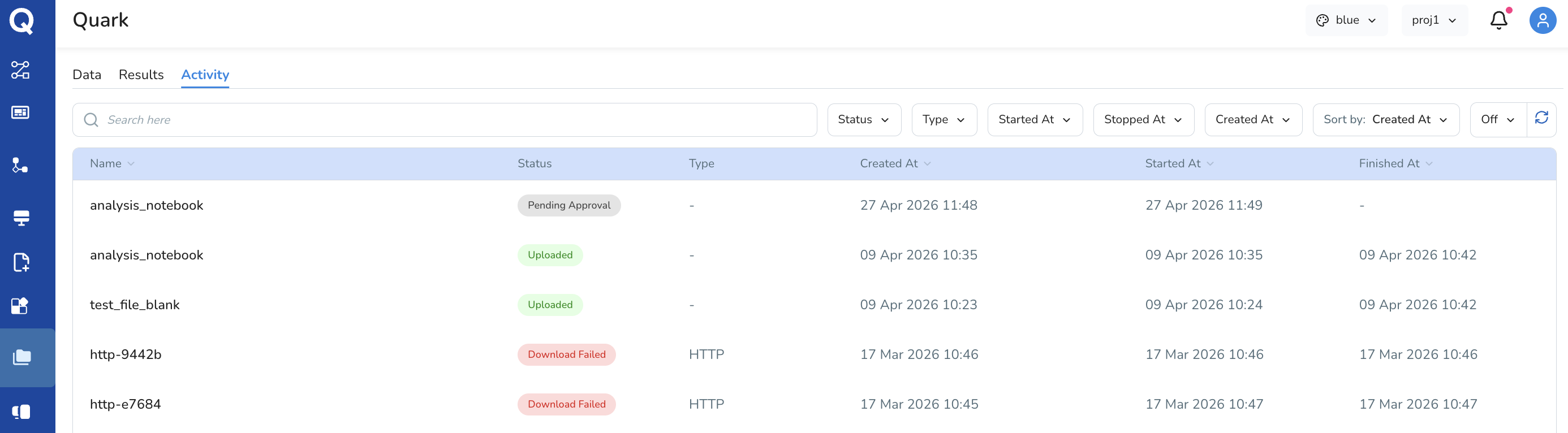
Activity Tab
The Activity tab provides a chronological log of file-level actions taken within your My Files workspace. This includes events such as file uploads, deletions, and any other modifications.
Activity logs are useful for:
- Auditing which files were uploaded and when
- Tracking changes made to datasets over time
-
Troubleshooting issues related to missing or unexpected files
Tips and Best Practices
-
Name files clearly before uploading. Pipeline run history is easier to follow when input file names reflect the sample, date, or experiment (e.g.
sample01_R1.fastq.gzrather thanfile1.fastq.gz). -
Check the Data tab before starting a run. All required input files should be uploaded and visible in the Data tab before you begin configuring a pipeline run.
-
Results are tied to runs. If you need to re-run a pipeline with different parameters, results from each run are stored separately and labelled by their run name — so use descriptive run names to keep things organised.
Further Reading
- MyFiles CLI - transferring large files with the MyFiles CLI tool.
- Apps and Visualisation — mounting data files and results directories in visualisation apps